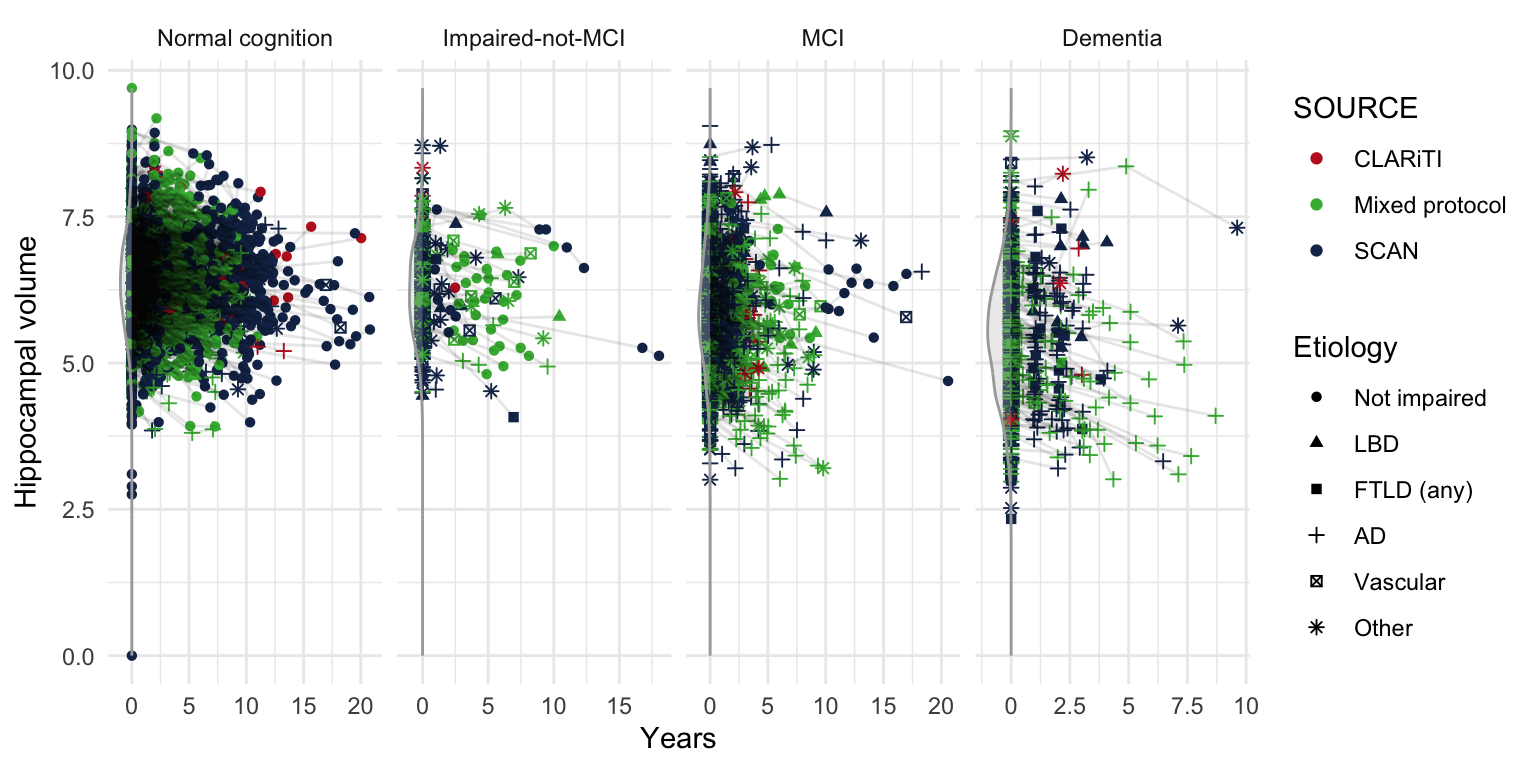
## Combine hippocampal volume from SCAN and Mixed Protocol (UDS) ----
mri_hipp <- mrisbm %>% # SCAN volumes
select(SOURCE, NACCID, DATE = SCANDT, HIPPOCAMPUS, ICV = CEREBRUMTCV) %>%
bind_rows(uds_mri %>% # Mixed Protocol (UDS) volumes
select(NACCID, MRIYR, MRIMO, MRIDY, HIPPOCAMPUS = HIPPOVOL, ICV = NACCICV) %>%
filter(HIPPOCAMPUS != 88.8888, ICV != 9999.999, !is.na(HIPPOCAMPUS)) %>%
mutate(
DATE = as.IDate(paste(MRIYR, MRIMO, MRIDY, sep='-')),
SOURCE = 'Mixed protocol')) %>%
distinct(NACCID, DATE, .keep_all = TRUE)
NACCETPR_levels <- names(rev(sort(table(uds_ftldlbd$NACCETPR))))
## Combine hipp volume, tau PET, amyloid PET, cognition, and demographics ----
dd_atn <- clariti_edc %>%
select(NACCID) %>% distinct() %>% mutate(`CLARiTI EDC` = 'Yes') %>%
full_join(mri_hipp %>% # Hippocampal volumes
select(NACCID, MRI_SOURCE = SOURCE, DATE, HIPPOCAMPUS, ICV),
by = 'NACCID') %>%
full_join(taupetnpdka %>%
arrange(NACCID, SCANDATE, desc(PROCESSDATE)) %>%
distinct(NACCID, SCANDATE, .keep_all = TRUE) %>%
select(NACCID, DATE = SCANDATE, Tau_PET = META_TEMPORAL_SUVR,
CTX_ENTORHINAL_SUVR, Tau_TRACER = TRACER, Tau_SOURCE = SOURCE) %>%
distinct(NACCID, DATE, .keep_all = TRUE),
by = c('NACCID', 'DATE')) %>%
full_join(amyloidpetgaain %>% # Amyloid PET
arrange(NACCID, SCANDATE, desc(PROCESSDATE)) %>%
distinct(NACCID, SCANDATE, .keep_all = TRUE) %>%
select(NACCID, DATE = SCANDATE, AMYLOID_STATUS, CENTILOIDS,
Amyloid_TRACER = TRACER, Amyloid_SOURCE = SOURCE) %>%
distinct(NACCID, DATE, .keep_all = TRUE),
by = c('NACCID', 'DATE')) %>%
full_join(phc_cognition %>% # Harmonized cognition
select(NACCID, NACCVNUM, PHC_MEM, PHC_EXF, PHC_LAN) %>%
distinct(NACCID, NACCVNUM, .keep_all = TRUE) %>%
left_join(uds_ftldlbd %>% # UDS visit dates
select(NACCID, NACCVNUM, DATE = VISITDATE),
by = c('NACCID', 'NACCVNUM')) %>%
distinct(NACCID, DATE, .keep_all = TRUE),
by = c('NACCID', 'DATE'))
stopifnot(with(dd_atn, !any(duplicated(paste(NACCID, DATE)))))
dd <- dd_atn %>%
full_join(uds_ftldlbd %>% # UDS diagnosis, etiology over time
filter(NACCID %in% c(dd_atn$NACCID, clariti_edc$NACCID)) %>%
mutate(
NACCETPR = factor(NACCETPR, levels = NACCETPR_levels)) %>%
select(NACCID, DATE = VISITDATE, NACCUDSD, NACCETPR, PARK) %>%
distinct(NACCID, DATE, .keep_all = TRUE),
by = c('NACCID', 'DATE')) %>%
left_join(uds_ftldlbd %>% # UDS demographics
mutate(BIRTHDATE = as.IDate(paste(BIRTHYR, BIRTHMO, 15, sep = '-'))) %>%
filter(!is.na(BIRTHDATE)) %>%
arrange(NACCID, NACCVNUM) %>%
select(NACCID, BIRTHDATE, RACE, SEX = NACCSEX, EDUC, HISPANIC = NACCHISP) %>%
group_by(NACCID) %>% # Carry information back/forward
tidyr::fill(.direction = "updown") %>% # to impute missing data
ungroup() %>%
filter(!duplicated(NACCID)), # One row of demographics per NACCID
by = 'NACCID') %>%
arrange(NACCID, DATE) %>%
group_by(NACCID) %>% # Carry Dx information forward/back
tidyr::fill(all_of(c("NACCUDSD", "NACCETPR", "PARK")), .direction = "downup") %>%
mutate(ICV = mean(ICV, na.rm = TRUE)) %>% # Average ICV over visits
ungroup() %>%
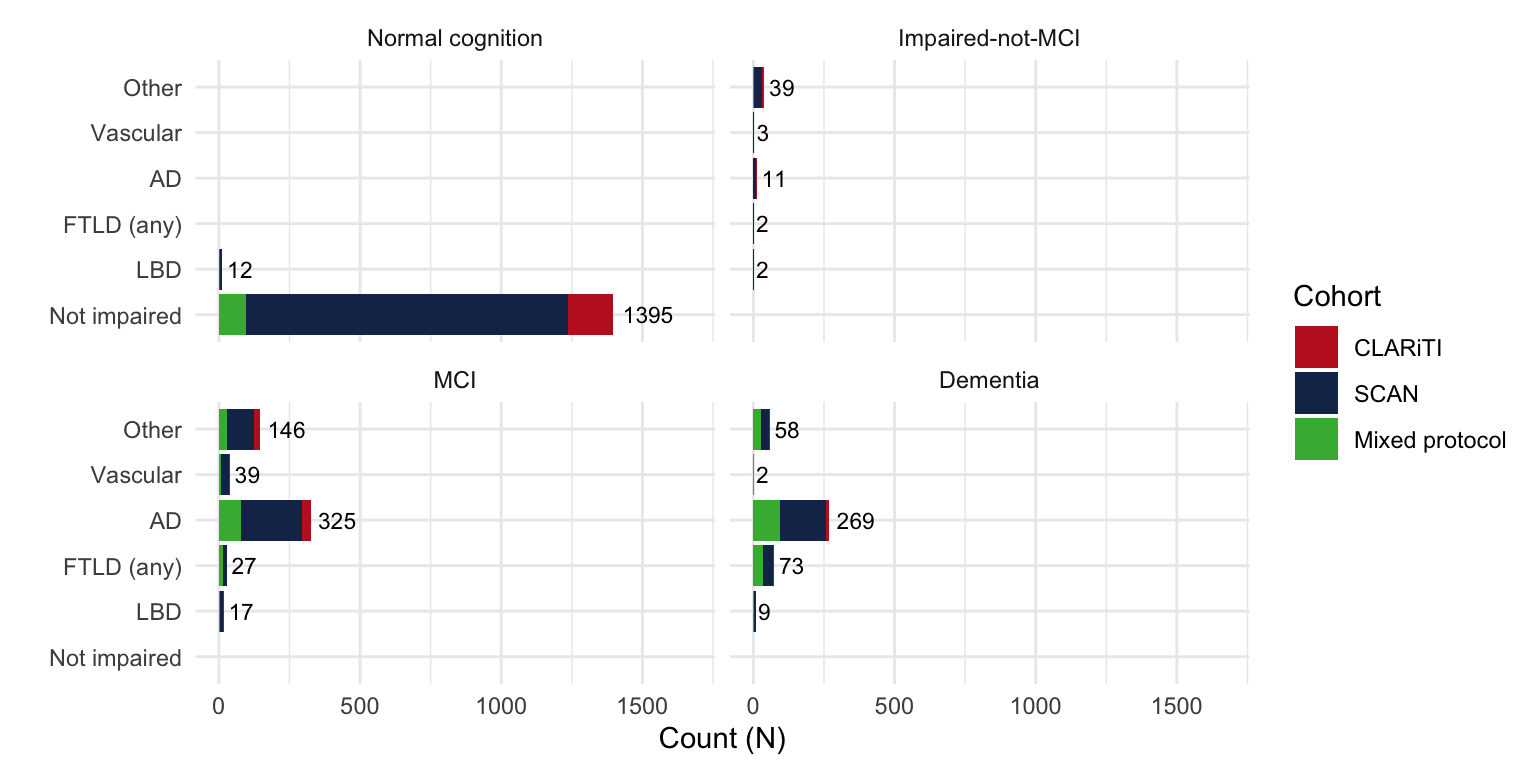
mutate(
Age = as.numeric(DATE - BIRTHDATE)/365.25,
Etiology = case_when( # Simplified/collapse diagnosis
NACCETPR == "Alzheimer's disease (AD)" ~ 'AD',
NACCETPR == 'Lewy body disease (LBD)' | PARK == 'Yes' ~ 'LBD',
NACCETPR %in% c("FTLD, other", "FTLD with motor neuron disease (e.g., ALS)") ~
"FTLD (any)",
NACCETPR == "Vascular brain injury or vascular dementia including stroke" ~ "Vascular",
NACCETPR == 'Not applicable, not cognitively impaired' ~ 'Not impaired',
TRUE ~ 'Other') %>%
factor(levels = c('Not impaired', 'LBD', "FTLD (any)", 'AD', "Vascular", 'Other')),
NACCUDSD = case_when(
is.na(NACCUDSD) ~ 'Unknown',
TRUE ~ NACCUDSD) %>%
factor(levels = c("Normal cognition", "Impaired-not-MCI",
"MCI", "Dementia", 'Unknown')))
stopifnot(with(dd, !any(duplicated(paste(NACCID, DATE)))))
CLARiTI_id <- dd %>%
filter(MRI_SOURCE == 'CLARiTI' | Tau_SOURCE == 'CLARiTI' | Amyloid_SOURCE == 'CLARiTI') %>%
pull(NACCID) %>% unique()
SCAN_id <- dd %>%
filter(MRI_SOURCE == 'SCAN' | Tau_SOURCE == 'SCAN' | Amyloid_SOURCE == 'SCAN') %>%
pull(NACCID) %>% unique() %>%
setdiff(CLARiTI_id)
Mixed_id <- setdiff(dd$NACCID, c(CLARiTI_id, SCAN_id))
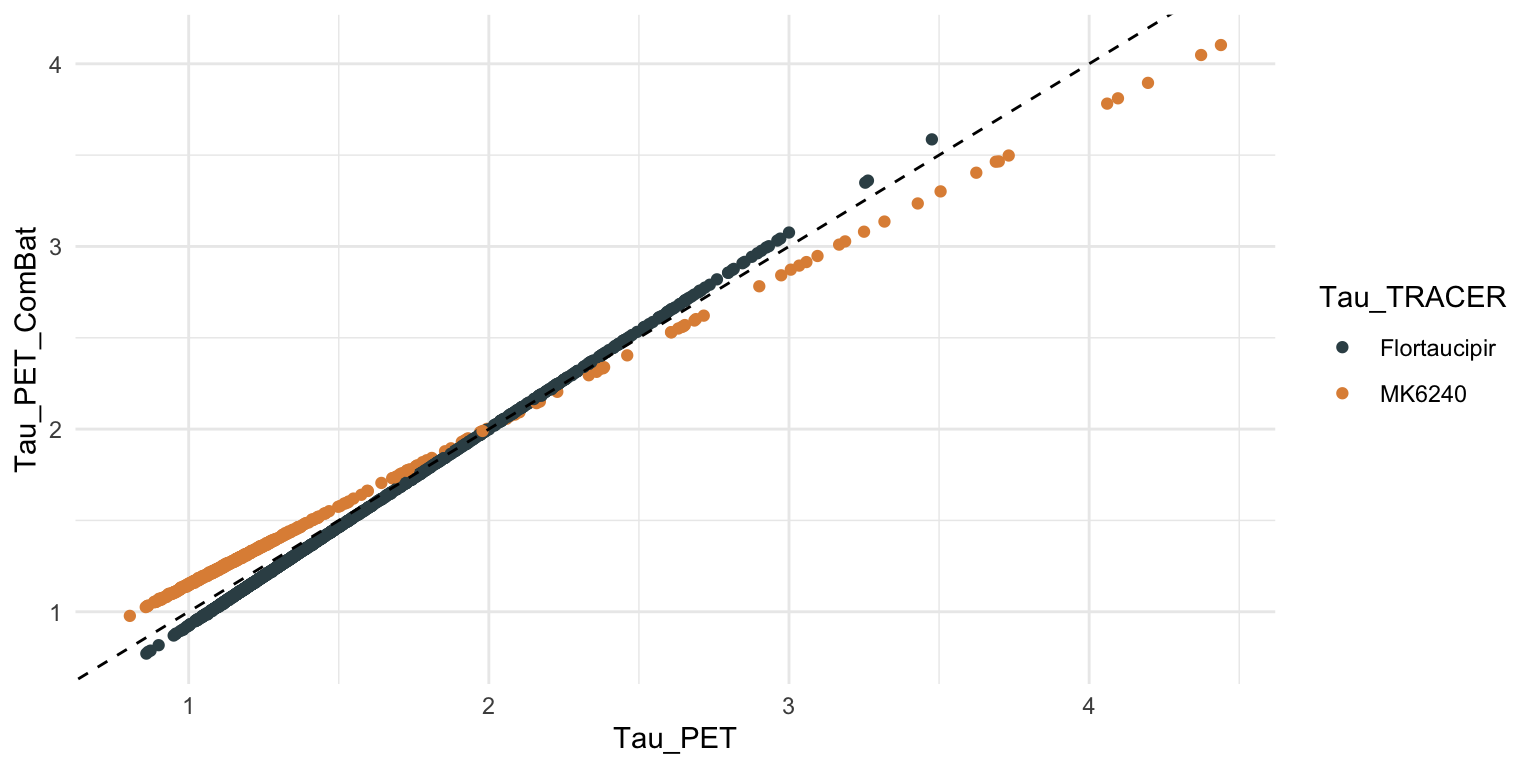
# harmonize tau PET data ----
tmp <- dd %>% filter(!is.na(Tau_PET) & !is.na(Age))
tmp$Tau_PET_ComBat <- sva::ComBat(
dat=t(tmp[, c('Tau_PET', 'CTX_ENTORHINAL_SUVR')]),
batch=tmp$Tau_TRACER,
mod=model.matrix(~-1 + Age, tmp), par.prior=TRUE, prior.plots=FALSE)[1,]
dd <- dd %>%
left_join(tmp %>% select(NACCID, DATE, Tau_PET_ComBat),
by = c('NACCID', 'DATE'))
# x-sectional data ----
dd_cross <- dd %>%
group_by(NACCID) %>%
fill(everything(), .direction = "updown") %>%
filter(!duplicated(NACCID)) %>%
ungroup() %>%
mutate( # add labels for tables
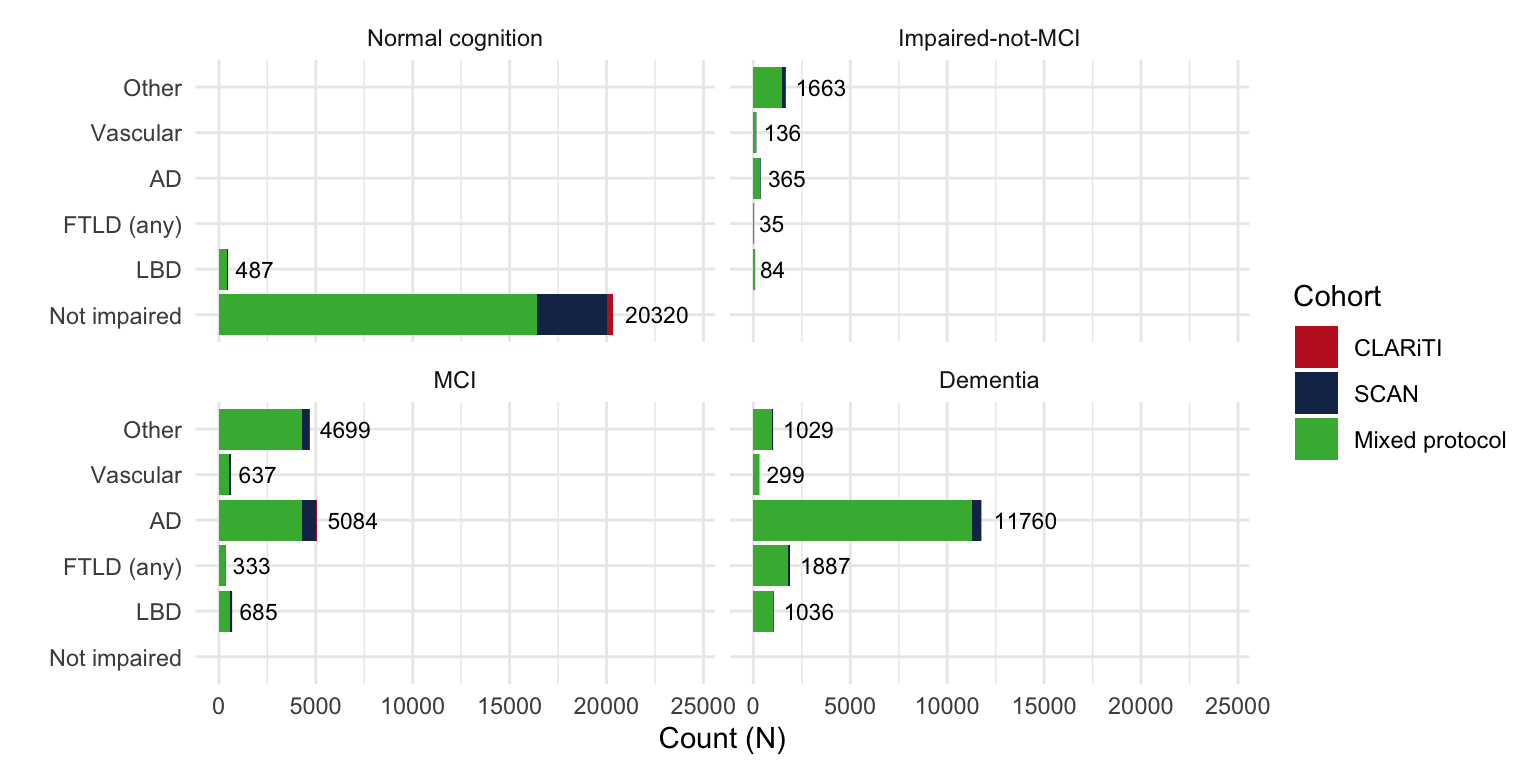
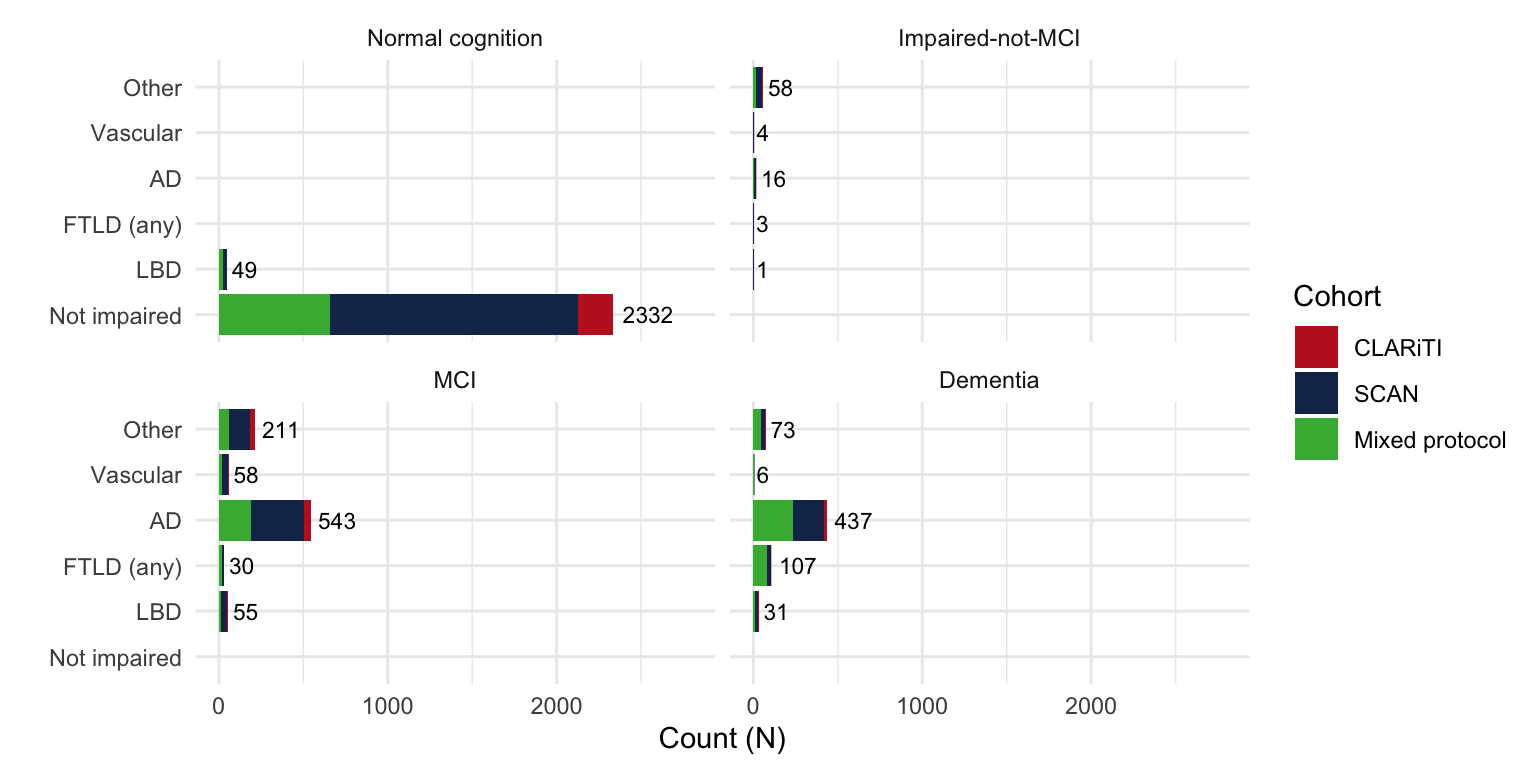
Cohort = case_when(
NACCID %in% clariti_edc$NACCID ~ 'CLARiTI',
NACCID %in% CLARiTI_id ~ 'CLARiTI',
NACCID %in% SCAN_id ~ 'SCAN',
TRUE ~ 'Mixed protocol') %>%
factor(levels = c('CLARiTI', 'SCAN', 'Mixed protocol')),
PHC_MEM = structure(PHC_MEM, label = 'Harmonized Memory'),
PHC_EXF = structure(PHC_EXF, label = 'Harmonized Exec Function'),
PHC_LAN = structure(PHC_LAN, label = 'Harmonized Language'),
EDUC = structure(EDUC, label = 'Education (yrs)'),
Age = structure(Age, label = 'Age (yrs)'),
SEX = structure(SEX, label = 'Sex'),
RACE = structure(RACE, label = 'Race'),
HISPANIC = structure(HISPANIC, label = 'Hispanic'),
AMYLOID_STATUS = structure(AMYLOID_STATUS, label = 'Amyloid status'),
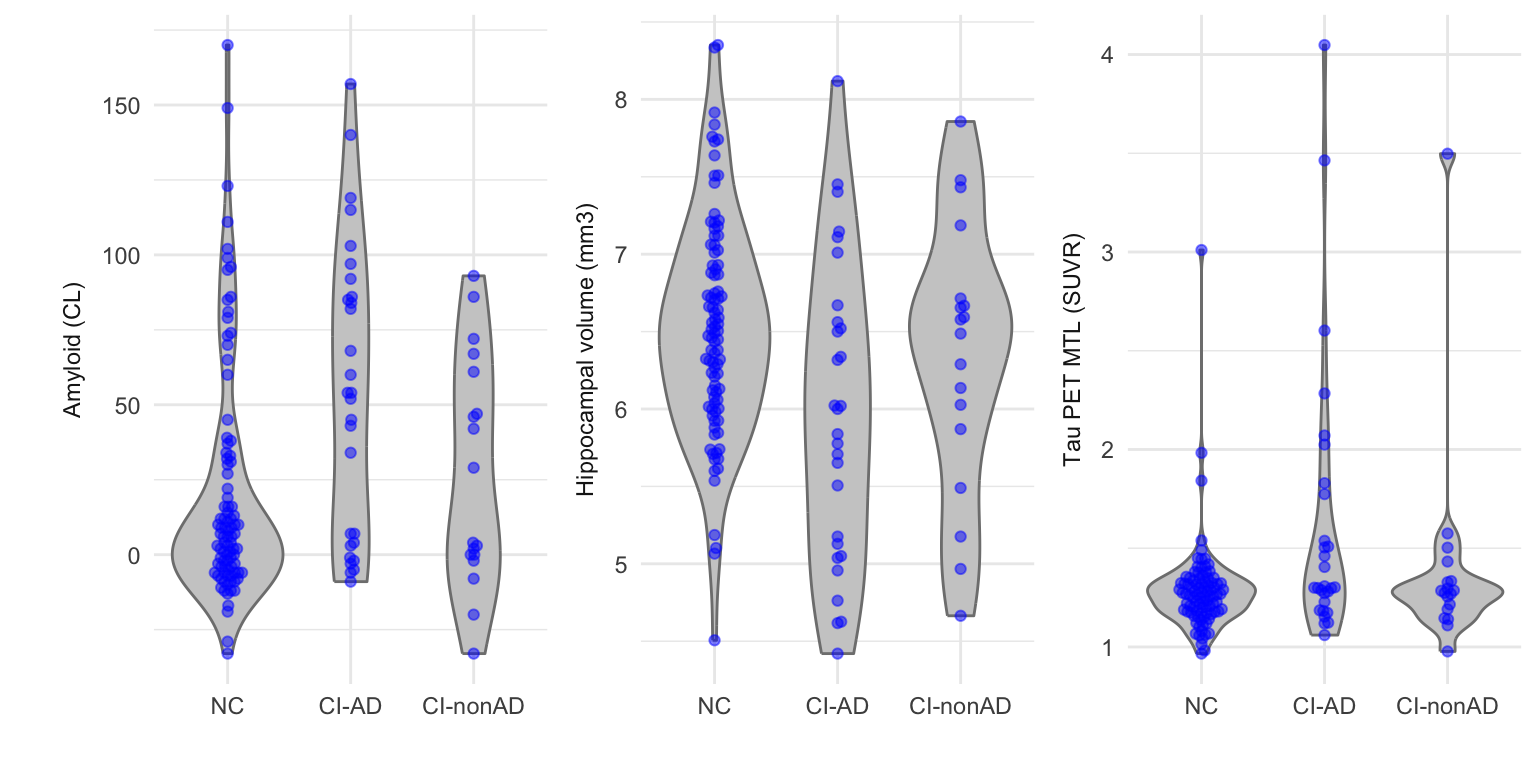
CENTILOIDS = structure(CENTILOIDS, label = 'Amyloid (CL)'),
HIPPOCAMPUS = structure(HIPPOCAMPUS, label = 'Hippocampal volume (mm3)'),
Tau_PET = structure(Tau_PET, label = 'Tau PET (SUVR)'),
Tau_PET_ComBat = structure(Tau_PET_ComBat, label = 'Tau PET (SUVR)'),
Tau_TRACER = structure(Tau_TRACER, label = 'Tau PET tracer'),
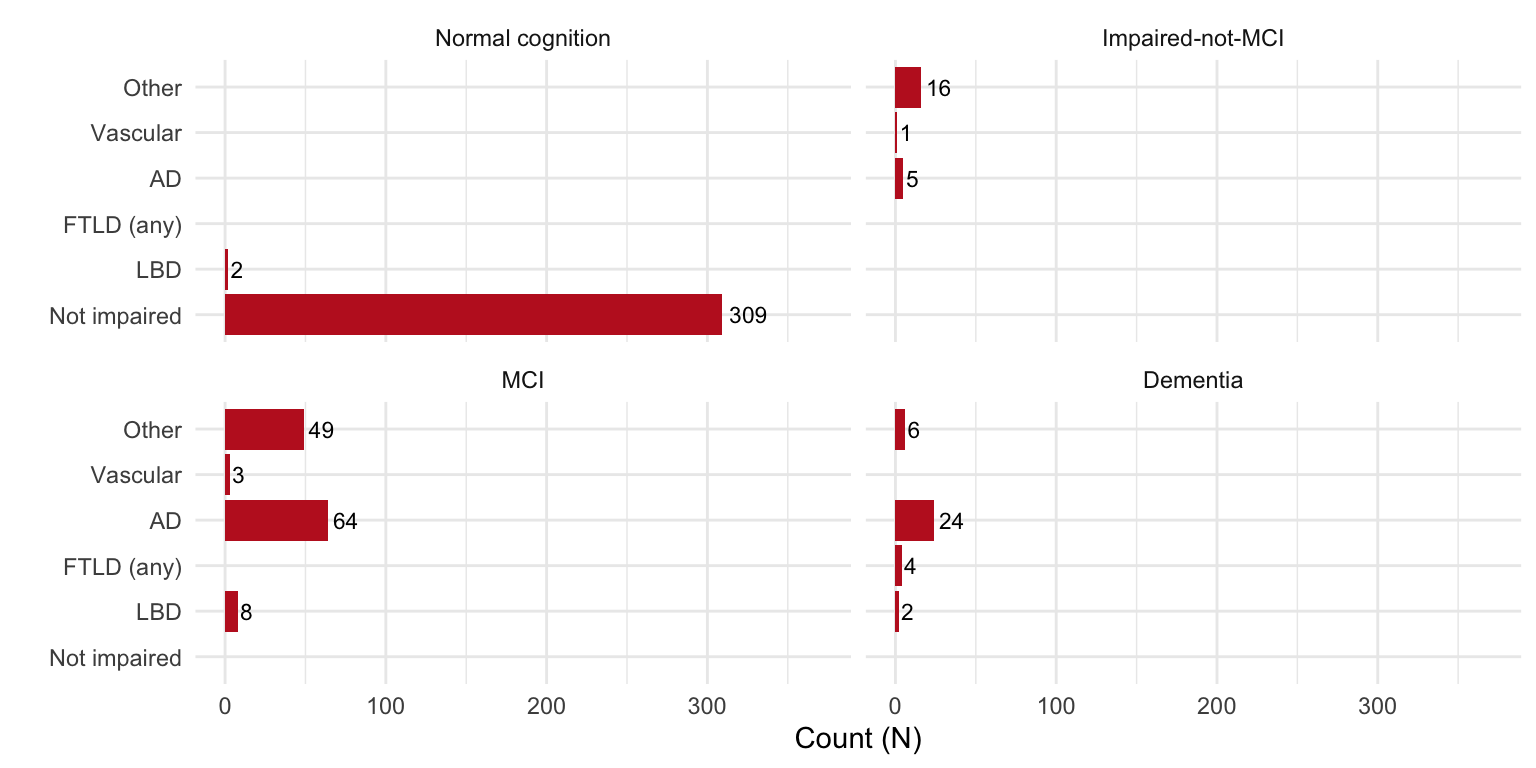
NACCETPR = structure(NACCETPR, label = 'Primary etiologic diagnosis'),
NACCUDSD = structure(NACCUDSD, label = 'Cognitive status at UDS visit'))